Analysis system developed for breast cancer diagnosis
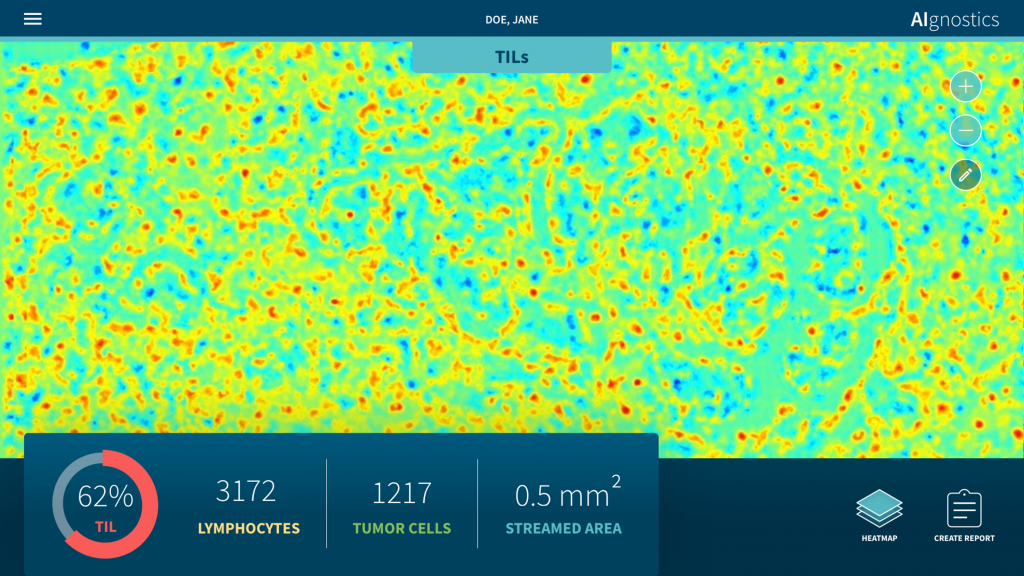
Researchers at Charité – Universitätsmedizin Berlin and TU Berlin have developed a new analysis system for breast cancer diagnosis based on tissue sections that uses artificial intelligence (AI). Two further developments make the system unique: Firstly, it integrates morphological, molecular and histological data in one evaluation for the first time. Secondly, it provides an explanation of the AI decision-making process in the form of heat maps. This enables doctors to understand the result of the AI analysis and check it for plausibility. Artificial intelligence thus becomes explainable – a decisive and indispensable step forward if AI systems are to be used in the future to support medicine in everyday clinical practice. The research results have now been published in Nature Machine Intelligence*.
Cancer medicine is increasingly concerned with the molecular characterisation of tumour tissue samples. Among other things, the methylation state of the DNA, gene expression, somatic mutations or protein expression in the pathological specimens are determined. At the same time, the realisation is gaining ground that cancer progression is closely related to the connection of cancer cells to each other and the interaction with the surrounding tissue – including the immune system. While microscopic techniques allow the investigation of biological processes with high spatial resolution, molecular markers can only be collected microscopically to a limited extent. Instead, they are determined using proteins or DNA extracted from tissue samples. As a result, they usually do not allow spatial resolution, and therefore their relationship to the microscopic structures is typically unclear. An interdisciplinary research team has now been able to solve these problems with the help of AI.
“In breast cancer, it is known that the number of migrated immune cells, the so-called lymphocytes, in the tumour tissue has an influence on the prognosis of the patient. In addition, it is being discussed whether this number also has a predictive value – that is, whether it allows statements to be made about how well which therapy works,” says Prof. Dr. Frederick Klauschen from the Institute of Pathology at Charité.
“The problem is that we have good and reliable molecular data and good, spatially highly resolved histological data. But the crucial bridge between the imaging data and the high-dimensional molecular data has been missing until now,” adds Prof. Dr Klaus-Robert Müller, Professor of Machine Learning at TU Berlin. The two scientists have already been cooperating for several years under the umbrella of the national AI competence centre Berlin Institute for the Foundations of Learning and Data (BIFOLD), which is based at TU Berlin.
In the approach now published, precisely this symbiosis succeeded. “Our system enables the robust detection of pathological changes in microscopic images. In parallel, we provide a precise heat map visualisation that shows which pixel on the microscopic image contributed to the diagnosis of the algorithm and to what extent,” explains Prof. Müller. In addition, the researchers have taken the method one big step further: “Our analysis system has been trained with the help of machine learning methods so that it can also predict various molecular features, such as DNA methylation, gene expression or even protein expression in certain areas of the tissue from the histological images.”
Next on the agenda are certification and further clinical validations – including tests in routine pathological diagnostics. But Prof. Klauschen is convinced: “The method we have developed will allow histopathological tumour diagnostics to be more precise, standardised and thus qualitatively better in the future.
*Binder A et al. Morphological and molecular breast cancer profiling through explainable machine learning. Nat Mach Intell (2021), doi: 10.1038/s42256-021-00303-4
Links:
