Charité researchers simulate corona infection in human lungs and organoids
Berlin, 29.06.2022
A Berlin research group led by Charité – Universitätsmedizin Berlin has succeeded in simulating SARS-CoV-2 infection in human lungs, thus generating key insights into the infection mechanism. Using living lung samples cultivated in the laboratory, it shows that the COVID-19 pathogen is only able to directly infect the cells of the human alveoli to a very limited extent. In contrast, the majority of the viruses that enter the lungs are directly taken up by macrophages – cells of the innate immune defence – and trigger a targeted immune activation in them. The results have now been published in the European Respiratory Journal*.
Scientists around the world are still working to better understand the mechanism behind COVID-19 infection and the pneumonia and lung damage that sometimes accompanies it. Researchers from the Charité, the Berlin Institute of Health at the Charité (BIH), the Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC), the Robert Koch Institute and the Free University of Berlin have now analysed the replication and immune activation of SARS-CoV-2 viruses in human lungs. They have specifically focused on the cells of the human alveoli, also called alveoli, as well as the alveolar macrophages. These scavenger cells of our innate immune system destroy foreign particles, including infectious agents such as viruses and bacteria, and thus ensure the cleansing of the lungs.
Under the leadership of Prof. Dr. Andreas Hocke from the Medical Clinic with a focus on Infectiology and Pneumology at the Charité, the research team has found that SARS-CoV-2 infects only very few epithelial cells that line the surface of the alveoli and thus causes only very minor, direct tissue damage. This represents a decisive difference to MERS coronaviruses or influenza viruses, for example. At the same time, the scientists were able to prove that the ACE2 receptor required for SARS-CoV-2, which serves as an entry portal for the viruses, is detectable in only very few alveolar epithelial cells. This was revealed by extensive analyses using spectral microscopy.
“We were able to show the direct dependence of SARS-CoV-2 on its receptor in human lungs as well as in lung organoids – which are models of human alveoli that we obtained from stem cells of lung tissue – and thus exclude other, alternative receptors,” explains the study’s first author Dr. Katja Hönzke from the Medical Clinic with a focus on infectiology and pneumology. According to the study, if large amounts of viruses from the upper respiratory tract enter the alveoli, they do not multiply to a high degree in the resident epithelial cells of the lung, as is often the case with other severe viral infections, but are taken up directly by the phagocytes. “Using detailed bioinformatic analyses as well as autopsy tissue from people who died of COVID-19, we saw that the phagocytes change as a result of the uptake of the coronaviruses,” says the second first author of the study Dr. Benedikt Obermayer-Wasserscheid from BIH. These transformations in turn trigger different reactions in the context of pneumonia: The phagocytes release inflammatory messengers and can sometimes trigger very strong inflammatory cascades. The researchers also observed that the virus does not multiply in the immune cells.
Prof. Hocke classifies the results: “Our study suggests that severe lung damage in COVID-19 is due to immune activation triggered by macrophages rather than direct destruction of the alveoli by the virus. Thus, it contributes significantly to the understanding of the development of COVID-19 in the early phase of possible pneumonia and shows why SARS-CoV-2, in contrast to MERS coronaviruses, has a rather moderate course in the majority of cases.” It can therefore be assumed that the local immune mechanisms in the respiratory tissue eliminate the SARS-CoV-2 viruses very efficiently in the vast majority of cases and limit the inflammatory response. If this does not happen, which may be influenced by individual risk factors, severe and fatal courses can develop in rare cases. Prof. Hocke continues: “The lung models we have used are an excellent example of how alternatives to animal models based on human cells can be used, especially in research into zoonotic diseases. We have achieved this in close collaboration with Charité 3R, our facility for developing alternatives to animal testing.”
Subsequent work will now focus on studies of patient-specific organoid models in order to analyse in greater depth the influence of general risk factors such as age, gender, concomitant diseases and other medications on the activation of the inflammatory response. This knowledge could then be used to identify possible therapeutic approaches targeting the immune system.
*Hönzke K, Obermayer B, Mache C, et al. Human lungs show limited permissiveness for SARS-CoV-2 due to scarce ACE2 levels but virus-induced expansion of inflammatory macrophages. European Respiratory Journal (2022). doi: 10.1183/13993003.02725-2021
About the study
The work was funded by the German Research Foundation (DFG) in the Collaborative Research Centre “Innate Immunity of the Lung” (SFB-TR84), the Einstein Foundation Berlin (Einstein Centre 3R) and by the Federal Ministry of Education and Research (BMBF) as part of the joint project “Organ-specific stratification in COVID-19” (Organo-Strat). The joint project is part of the Network University Medicine (NUM), which was initiated and is coordinated by the Charité. The NUM unites the forces of the 36 university hospitals in Germany.
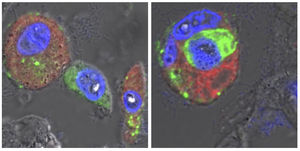
Caption: Microscopic images of lung tissue from a person who died of COVID-19: Alveolar macrophages (red, cell nuclei in blue) have taken up SARS-CoV-2 viral material (green). Between these scavenger cells is an epithelial cell of the human alveoli, which was detached from the tissue association by the infection (left image). The right image illustrates how these dead cells are taken up and disposed of by alveolar macrophages, alongside the viral material. © Charité I Andreas Hocke
Links:
Medizinische Klinik mit Schwerpunkt Infektiologie und Pneumologie
